PEG Evidence Categories
🌟 Why do we need PEG evidence categories?
Predicted Effector Gene (PEG) lists are built from diverse sources of evidence, ranging from genetic associations and computational predictions to functional assays and literature curation. Each source has its own origin, methodology, and level of confidence.
To ensure clarity and interoperability, PEGASUS group these diverse evidence types into standardised categories that are widely recognised in effector gene exploration.
➡️ Organising evidence into categories helps users:
- Interpret consistently – understand the general type of support behind each gene.
- Compare across studies – align evidence types, even when detailed sources differ.
- Build trust and transparency – see at a glance how predictions were made.
PEG Evidence Categories
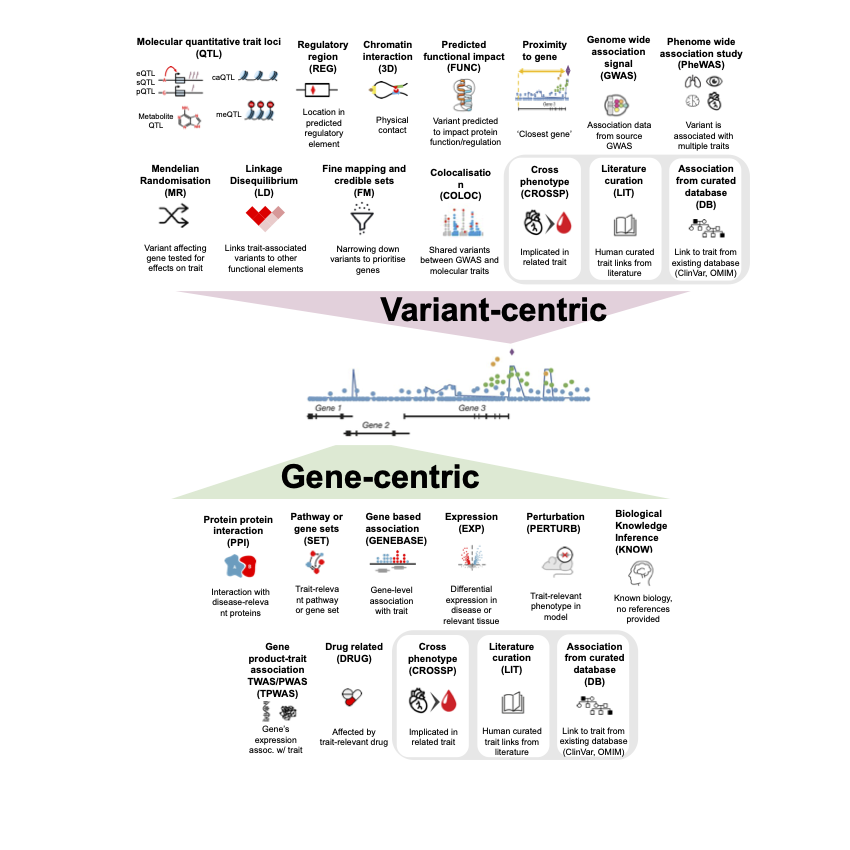
🔀 Variant-centric 🔀
| Evidence categories | Abbreviation | Explanation |
|---|---|---|
| Linkage disequilibrium | LD | Assessment of whether variant is correlated with another variant of interest and may act as a proxy. |
| Finemapping and credible sets | FM | Finemapping results - probability of variant being causal within a credible set, using Bayesian or probabilistic models |
| Colocalisation | COLOC | Variant affects two traits (typically a complex trait and a molecular phenotype) at the same locus. |
| Molecular QTL | QTL | Variant affects a molecular phenotype, e.g. gene expression (eQTL), splicing (sQTL), or protein expression (pQTL). |
| Mendelian Randomization (MR) | MR | Uses genetic variants as proxies for exposures to test their causal effect on outcomes. |
| Regulatory region | REG | Variant lies in open chromatin or enhancer/promoter elements in relevant tissue (e.g. ATAC-seq, DNase-seq, or histone mark data) |
| Chromatin interaction | CHROMATIN | Variant lies in a region physically interacting with a gene promoter via 3D chromatin architecture (e.g. Hi-C, Capture-C data). |
| Predicted functional impact | FUNC | Variant predicted to disrupt gene/protein function or regulatory motifs, e.g. via SIFT, PolyPhen, CADD. |
| Proximity to gene (distance) | PROX | Assessment of whether variant is within or near gene boundaries. |
| Genome-wide association (GWAS) signal | GWAS | P-value from source GWAS for association of variant with trait specified in metadata file |
| PheWAS (Phenome-Wide Association Study) | PHEWAS | Variant is associated with multiple traits, suggesting pleiotropic effects |
🧬 Gene-centric 🧬
| Evidence categories | Abbreviation | Explanation |
|---|---|---|
| Protein–protein interaction | PPI | Gene’s protein interacts with other disease-relevant proteins. |
| Pathway or gene sets | SET | Gene is part of a known pathway or complex relevant to the phenotype, e.g. results of enrichment analyses using Reactome or KEGG. |
| Gene-based association | GENEBASE | Aggregated analysis of association of variants in gene with trait (e.g. SKAT, MAGMA, burden tests) . |
| Expression | EXP | Gene is differentially expressed in relevant tissue or disease e.g. the gene is more highly expressed in phenotype-related tissues compared to others. |
| Perturbation | PERTURB | Gene perturbation causes phenotype-relevant effects in lab or model organisms (knock out animal/cell line, human organoid). |
| Biological Knowledge Inference | KNOW | Gene–phenotype relationships can be inferred based on known biology, without providing specific references or direct experimental evidence linking the specific gene to the phenotype. |
| Genetically predicted trait association (TWAS/PWAS) | TPWAS | Evidence from transcriptome- or proteome-wide association studies showing that gene’s genetically predicted expression or protein level is associated with phenotype, |
| Drug related | DRUG | Evidence from drug mechanism of action, e.g. gene encodes a known drug target or interacts with targets of drugs used to treat the phenotype, supporting therapeutic relevance. |
Variant or Gene centric evidence
| Evidence categories | Abbreviation | Explanation |
|---|---|---|
| Cross-phenotype | CROSSP | Gene or variant already established in a related phenotype (biologically similar). |
| Literature curation | LIT | Human-curated gene or variant–disease links from literature. |
| Association from curated database | DB | Variant or Gene is curated as causal or related to the phenotype from existing database, like ClinVar, ClinGen, OMIM, etc. |